summary
XGBoost: A Scalable Tree Boosting System
Presented by
Jiang, Cong
Song, Ziwei
Ye, Zhaoshan
Zhang, Wenling
Introduction
The Extreme Gradiant Boosting, also known as XGBoost, has been the leading model that dominates Kaggle, as well as other machine learning and data mining challenges. The main reason to its success is that this tree boosting system is highly scalable , which means it could be used to solve various problems with significantly less time and fewer resources.
This paper is written by the father of XGBoost, Tianqi Chen. It gives us an overview on how he used algorithmic optimizations as well as some important systems to develop XGBoost. He explained it in the following manner:
1. Based on gradient tree boosting algorithm, Chen makes some modifications to the regularized objective function and introduces an end-to-end tree boosting system. He also applies shrinkage and feature subsampling to prevent overfitting,
2. The system supports existing split finding algorithms including exact greedy and approximate algorithms. For weighted data, Chen proposes a distributed weighted quantile sketch algorithm which is provable; while for sparse data, he develops a novel sparsity-aware algorithm which is much faster than the naive model.
3. Storing the data in blocks is Chen's method to reduce sorting cost. The cache-aware block structure is highly effective for out-of-core tree learning.
4. XGBoost is evaluated by designed experiments with 4 datasets. Chen compares it with adjusted setting as well as other systems to test its performance.
Tree Boosting
Regularized Learning Objective
Suppose the given data set has m features and n examples. A tree ensemble model uses K additive functions to predict the output.
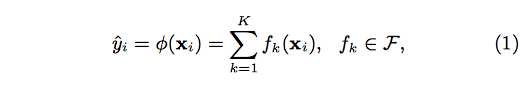
where F is the space of regression trees(CART). The author suggests the following regularized objective function to make improvements comparing to gradient boosting
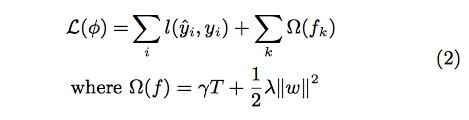
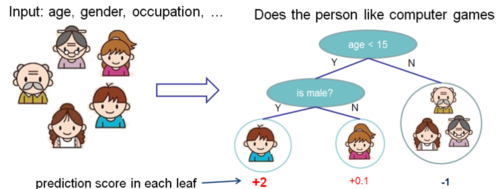
Regression tree contains continuous scores on each leaf, which is different with decision trees we have learned in class. The following 2 figures represent how to use the decision rules in the trees to classify it into the leaves and calculate the final prediction by summing up the score in the corresponding leaves.
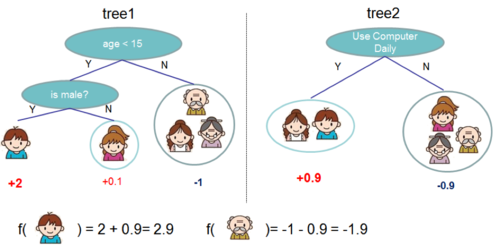
Gradient Tree Boosting
The tree ensemble model (Equation 2.) is trained in an additive manner. We need to add a new [math]\displaystyle{ f }[/math] to our objective.
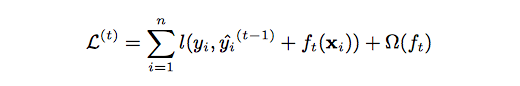
After applying the second-order Taylor expansion to the above formula and removing constant terms, we can finally have
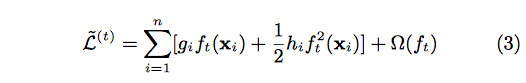
Define [math]\displaystyle{ I_{j} = \{i| q(X_{i}) \} }[/math] As the instance set of leaf j. We can rewrite Equation (3) as
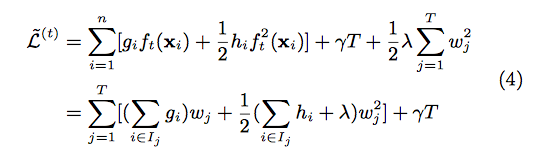
By computing the optimal weight and corresponding optimal value, we can use following equation as the scoring function which can measure the quality of a tree structure. Lower score means better tree structure.
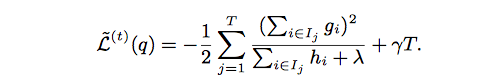
The following figure shows an example how this score is calculated.
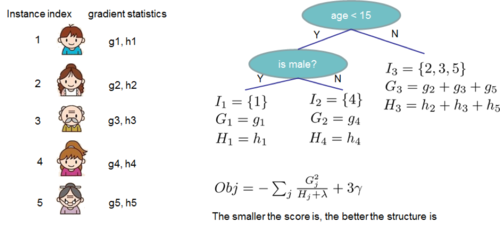
Normally it is impossible to enumerate all the possible tree structures. Assume [math]\displaystyle{ I_{L} }[/math] and [math]\displaystyle{ I_{R} }[/math] are the instance sets of left and right nodes after the split, and [math]\displaystyle{ I = I_{L} + I_{R} }[/math]. Then the loss reduction after the split is given by
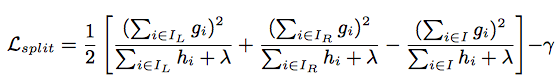
Shrinkage and Column Subsampling
In addition to the regularized objective, the paper also introduces two techniques to prevent over-fitting.
Method 1: Shrinkage. After each step of tree boosting, we scale newly added weights by a shrinkage factor η, which reduces the influence of every individual tree and leaves the space for future trees.
Method 2: Column (feature) subsampling. We try to build each individual tree with only a portion of predictors chosen randomly. This procedure encourages the variance between the trees (i.e. no same tree occurs). Implemented in an opensource packages, this method prevents over-fitting, compared with the traditional row subsampling, and speeds up the computations of parallel algorithm as well.
Split Finding Algorithms
The ways to find the best split can be realized by two main methods.
Basic exact greedy algorithm: enumerate over all the possible splits on all the features, which is computationally demanding. Sort the data first, and visit the data to accumulate the gradient statistics greedily.
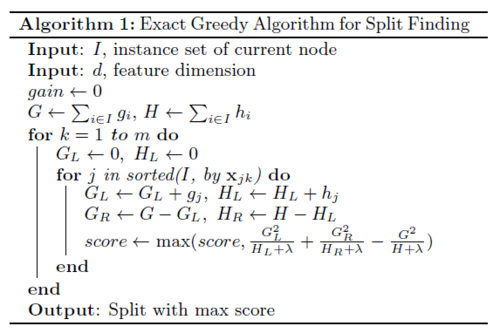
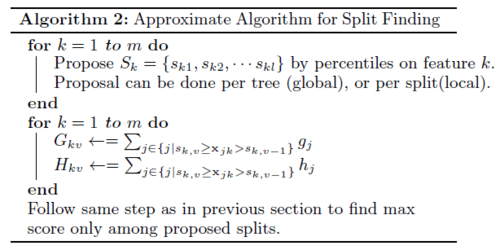


Approximate algorithm: To achieve an effective gradient tree boosting model, The author introduces an approximate algorithm. The algorithm first proposes candidate splitting points by the percentiles of feature distribution, maps the continuous features into buckets split and then finds the best solution among the proposals by the aggregated statistics. The algorithm can be varied by the time the proposal is given. The global variant: all candidate splits are proposed at the initial phase of tree construction and are used for all split finding. It requires fewer proposal steps. The local variant: the candidate splits are refined after each split. It requires fewer candidate points, so it is potentially more appropriate for deeper trees. Methods can be chosen according to users’ needs.
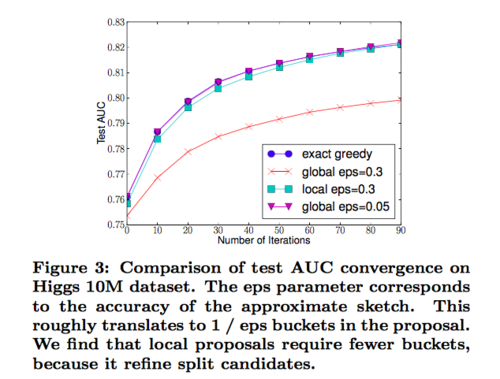
Weighted Quantile Sketch
Sparsity--aware Split Finding
In real world problem, it is quite common for the input X to be sparse. The reasons that causes for sparsity are following: 1. presence of missing values in the data; 2.frequent zero entries in the statistics; and, 3.artifacts of feature engineering such as one-hot encoding.
Evaluations
System Implementation
Implemented as an open source package, XGBoost is flexible, portable and reusable. It also supports various languages and ecosystems.
Dataset and Setup
Table 2 summarizes the 4 datasets used in the experiments.
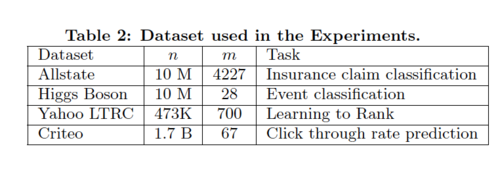
The first three datasets are used for the single machine parallel setting, which is run on a Dell PowerEdge R420 with two 8-core Intel Xeon and memory 64GB. Meanwhile, AWS c3.8xlarge machine is used for experiment in distributed and the out-of-core setting, which uses the criteo dataset. All available cores in the machines are devoted into the experiments, unless otherwise specified.
Experiments
The classification experiment uses Higgs Boson dataset to classify whether an event corresponds to the Higgs boson. The goal is to evaluate XGBoost using Exact Greedy Methods and compare it with column sampling XGBoost and two other well-known systems. Table 3 below summarizes the results.
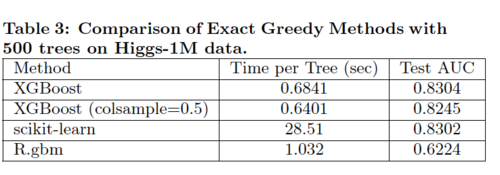
Overall, both XGBoost and scikit-learn performs better than R.gbm by 20% of Test AUC. In terms of time, however, XGBoost runs significantly faster than both scikit-learn and R.gbm. Note that XGBoost with column sampling performs slightly worse because the number of key features is limited. We could conclude that XGBoost using the exact greedy algorithm without column sampling works the best under this scenario.
The learning to rank experiment compares XGBoost(using exact greedy algorithm) with pGBRT(using approximate algorithm) in order to evaluate the system's performance on rank problem.
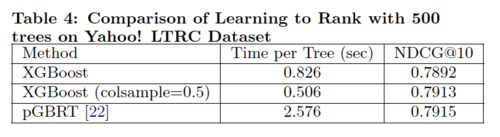
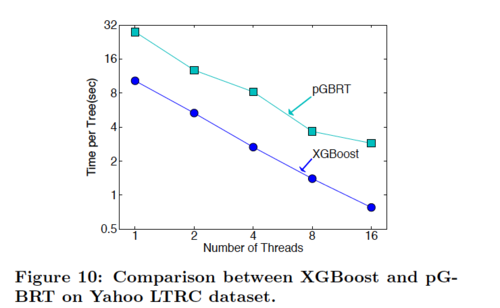
According to Table 4, XGBoost with column sampling performs almost as well as pGBRT, while using only 1/5 of the time. Note that pGBRT is the best previously published system to solve this rank problem.
The out-of-score and distributed experiments demonstrate XGBoost's ability to handle large data with a fast speed.
Conclusion
The writer, Tianqi Chen, proposed a scalable end-to-end tree boosting system with cache access patterns, block structure and other essential elements. He also introduced two novel split finding algorithms, one is sparsity-aware algorithm and the other is weighted quantile sketch with proofs. Compared to other popular systems, XGBoost gives state-of-the-art results in different experiment settings, which explains why it it considered one of the best and fastest algorithm for structured or tabular data.