a fair comparison of graph neural networks for graph classification
Presented By
Jaskirat Singh Bhatia
Background
Experimental reproducibility and replicability are critical topics in machine learning. Authors have often raised concerns about their lack in scientific publications to improve the quality of the field. Recently, the graph representation learning field has attracted the attention of a wide research community, which resulted in a large stream of works. As such, several Graph Neural Network models have been developed to effectively tackle graph classification. However, experimental procedures often lack rigorousness and are hardly reproducible. The authors tried to reproduce the results from such experiments to tackle the problem of ambiguity in experimental procedures and the impossibility of reproducing results. They also Standardized the experimental environment so that the results could be reproduced while using this environment.
Graph Neural Networks
A graph is a data structure consisting of nodes and edges. Graph neural networks are models which take graph-structured data as input, and capture information of the input graph, such as relation and interaction between nodes. In graph neural networks, nodes aggregate information from their neighbors. The key idea is to generate representations of nodes depending on the graph structure.
Graph neural networks can perform various tasks and have been used in many applications. Some simple and typical tasks include classifying the input graph or finding a missing edge/ node in the graph. One example of real applications where GNNs are used is social network prediction and recommendation, where the input data is naturally structural.
Problems in Papers
Some of the most common reproducibility problems encountered in this field concern hyperparameters selection and the correct usage of data splits for model selection versus model assessment. Moreover, the evaluation code is sometimes missing or incomplete, and experiments are not standardized across different works in terms of node and edge features.
These issues easily generate doubts and confusion among practitioners that need a fully transparent and reproducible experimental setting. As a matter of fact, the evaluation of a model goes through two different phases, namely model selection on the validation set and model assessment on the test set. Clearly, to fail in keeping these phases well separated could lead to over-optimistic and biased estimates of the true performance of a model, making it hard for other researchers to present competitive results without following the same ambiguous evaluation procedures.
Risk Assessment and Model Selection
Risk Assesment
The goal of risk assessment is to provide an estimate of the performance of a class of models. When a test set is not explicitly given, a common way to proceed is to use k-fold Cross Validation. As model selection is performed independently for each training/test split, they obtain different “best” hyper-parameter configurations; this is why they refer to the performance of a class of models.
Model Selection
The goal of model selection, or hyper-parameter tuning, is to choose among a set of candidate hyperparameter configurations the one that works best on a specific validation set.
Reproducibility Issues
The GNN's were selected based on the following criteria
1. Performances obtained with 10-fold CV
2. Peer reviews
3. Strong architectural differences
4. Popularity
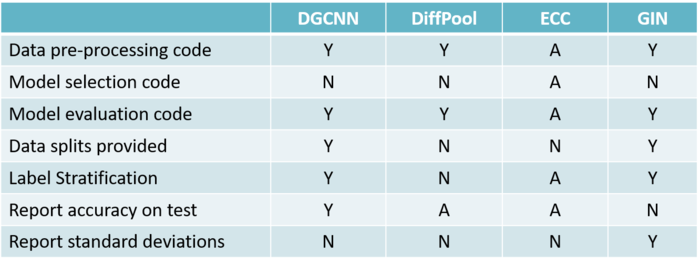
Criteria to assess the quality of evaluation and reproducibility was as follows
1. Code for data pre-processing
2. Code for model selection
3. Data splits are provided
4. Data is split by means of a stratification technique
5. Results of the 10-fold CV are reported correctly using standard deviations
Using the following criteria, 4 different papers were selected and their assessment on the quality of evaluation and reproducibility is as follows:
Experiments
They re-evaluate the above-mentioned models on 9 datasets (4 chemical, 5 social), using a model selection and assessment framework that closely follows the rigorous practices as described earlier. In addition, they implemented two baselines whose purpose is to understand the extent to which GNNs are able to exploit structural information.
Datasets
All graph datasets used are publicly available (Kersting et al., 2016) and represent a relevant subset of those most frequently used in literature to compare GNNs.
Features
In GNN literature, it is common practice to augment node descriptors with structural features. In general, good experimental practices suggest that all models should be consistently compared to the same input representations. This is why they re-evaluate all models using the same node features. In particular, they use one common setting for the chemical domain and two alternative settings as regards the social domain.
Baseline Model
They adopted two distinct baselines, one for chemical and one for social datasets. On all chemical datasets but for ENZYMES, they follow Ralaivola et al. (2005); Luzhnica et al. (2019) and implement the Molecular Fingerprint technique. On social domains and ENZYMES (due to the presence of additional features), they take inspiration from the work of Zaheer et al. (2017) to learn permutation-invariant functions over sets of nodes.
Experimental Setting
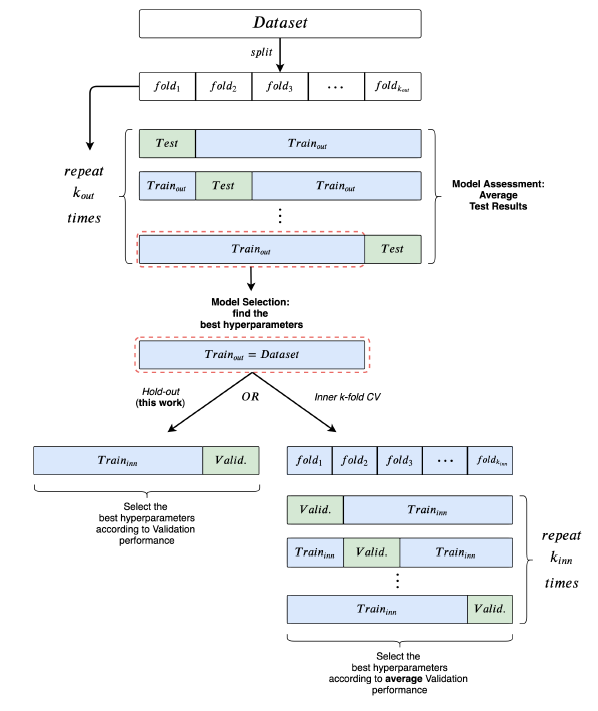
1. Used a 10-fold CV for model assessment and an inner holdout technique with a 90%/10% training/validation split for model selection.
2. After each model selection, they train three times on the whole training fold, holding out a random fraction (10%) of the data to perform early stopping.
3. The final test fold score is obtained as the mean of these three runs
4. To be consistent with literature, they implemented early stopping with patience parameter n, where training stops if n epochs have passed without improvement on the validation set.
Hyper-Parameters
1. Hyper-parameter tuning was performed via grid search.
2. They always included the hyper-parameters used by other authors in their respective papers.
Computational Considerations
As their research included a large number of training-testing cycles, they had to limit some of the models by:
1. For all models, grid sizes ranged from 32 to 72 possible configurations, depending on the number of hyper-parameters to choose from.
2. Limited the time to complete a single training to 72 hours.
Conclusion
1. Highlighted ambiguities in the experimental settings of different papers
2. Proposed a clear and reproducible procedure for future comparisons
3. Provided a complete re-evaluation of four GNNs
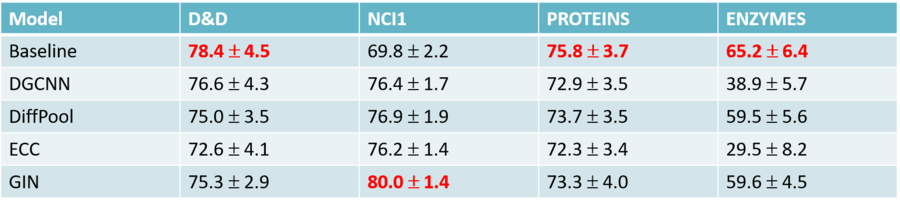
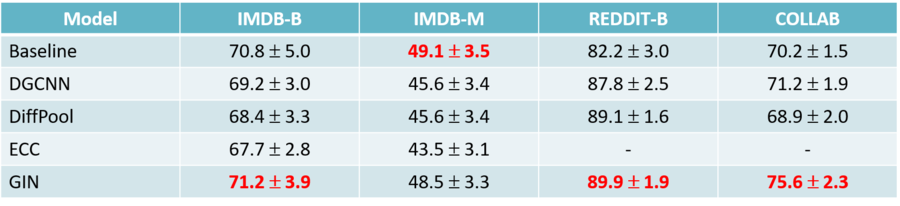
4. Found out that structure-agnostic baselines outperform GNNs on some chemical datasets, thus suggesting that structural properties have not been exploited yet.
References
- Davide Bacciu, Federico Errica, and Alessio Micheli. Contextual graph Markov model: A deep and generative approach to graph processing. In Proceedings of the International Conference on Machine Learning (ICML), volume 80 of Proceedings of Machine Learning Research, pp. 294–303. PMLR, 2018.
- Paul D Dobson and Andrew J Doig. Distinguishing enzyme structures from non-enzymes without alignments. Journal of molecular biology, 330(4):771–783, 2003.
- Will Hamilton, Zhitao Ying, and Jure Leskovec. Inductive representation learning on large graphs. In Advances in Neural Information Processing Systems (NIPS), pp. 1024–1034. Curran Associates, Inc., 2017.
- Kristian Kersting, Nils M. Kriege, Christopher Morris, Petra Mutzel, and Marion Neumann. Benchmark data sets for graph kernels, 2016. URL http://graphkernels.cs.tu-dortmund. de.