U-Time:A Fully Convolutional Network for Time Series Segmentation Applied to Sleep Staging Summary
Introduction
During sleep, the brain goes through different sleep stages, each characterised by brain and body activity patterns. Stages can be determined by measurements in a so called polysomnography study (PSG), which includes measurements of brain activity by EEG, eye movement and facial muscle activity. The process of mapping the transitions between sleep stages is called sleep staging and provides the basis for diagnosis of sleeping disorders. Traditionally, sleep staging is done manually by splitting the measurements of a PSG into 30 second segments, each containing multiple channels of data, and classifying the segments individually. Since this requires a lot of expertise and time, automatization is of interest. Fast and reliable automated sleep staging could help with diagnosis and help find novel biomarkers for disorders (Perslev et al., 2019).
State of the art sleep staging classifiers employ convolutional and recurrent layers. The problem with recurrent neural nets is that they can be difficult to tune and optimize and might need hyperparameter tuning to be suitable for different data sets. This means they are often specially trained to be applied on one dataset alone and might be difficult to use for non-experts in a more general setting (Perslev et al., 2019).
This paper introduces U-Time, a feed-forward convolutional network for sleep staging, which treats segmentation similar to how the popular image classifier U-net treats image segmentation. It does not need hyperparameter or architectural tuning to be applied to variable data sets, and it is able to classify sleep stages at any temporal resolution (Perslev et al., 2019).
Previous Work
Recently, there has been much interest in using machine learning techniques for analyzing physiological time series (Faust et al., 2018). Multiple neural network-based systems have been developed to classify different sleep-wake stages in humans, babies, and even cats (Claude et al., 1998). However, a drawback of recurrent neural networks is that they are difficult to tune and optimize in practice, resulting in many being replaced with feed-forward systems that don't lose accuracy (Bai et al., 2018; Chen & Wu, 2017; Vaswani et al., 2017). Here, U-Time is a feed-forward convolutional neural network that does not require hyperparameter or architectural tuning; in particular, it uses dilated convolutions to aggregate multi- scale contextual information without losing resolution or requiring the images to be rescaled (Yu & Koltun, 2015).
Methods
U-Time is a fully convolutional encoder-decoder network. Inspired from U-Net, U-time performs 1D time series segmentation by mapping a whole sequence to a dense segmentation in a single pass.
Consider x [math]\displaystyle{ \in \mathbb{R}^{T \times i\times C} }[/math] as T concatenated physiological signals of length i with C channels. U-time then makes predictions on the T physiological signal segments at once and directly maps the input to K confidence scores per segment. More explicitly, U-net maps x [math]\displaystyle{ \in \mathbb{R}^{T \times i\times C} \rightarrow \mathbb{R}^{T\times K} }[/math].
Architecture
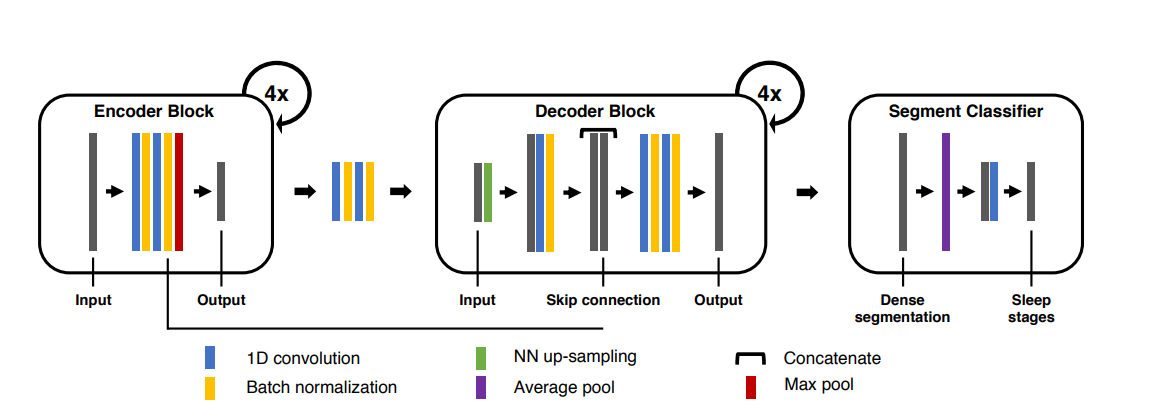
Encoder Block
The encoder consists of four convolutional blocks as shown above. All convolutional blocks preserve input dimensionality through zero padding. Each block in the encoder performs two consecutive convolutions with max pooling occurring afterward. Throughout the four blocks, pooling windows are 10, 8, 6, and 4 respectively. Dilated convolutional layers are also used in lieu of conventional convolutional layers. This aggressive down-sampling reduces both computational and memory requirements in addition to providing a very large receptive field. The maximum theoretical receptive field of U-time corresponds to approximately 5.5 minutes given a 100Hz signal.
Decoder Block
The decoder also consists of four convolutional blocks as shown above. Each block performs nearest neighbor up-sampling followed by conventional convolution with kernel sizes 4,6,9 and 10 and batch normalization. The resulting feature maps are then concatenated (along the filter dimension) with the corresponding feature maps computed by the encoder at the same scale. Then Two convolutional layers, both followed by batch normalization, process the concatenated feature maps in each block. Finally, a pointwise convolution with K filters results in K scores for each sample in the input sequence.
Segment Classifier
The output from the decoder block is then fed into the segment classifier. This serves as a trainable link between the dense segmentation and the final output classifications. This link uses an average pooling layer followed by a point-wise convolution to average out the scores from the dense segmentation into a sleep stage classification for each segment of physiological signals.
Results
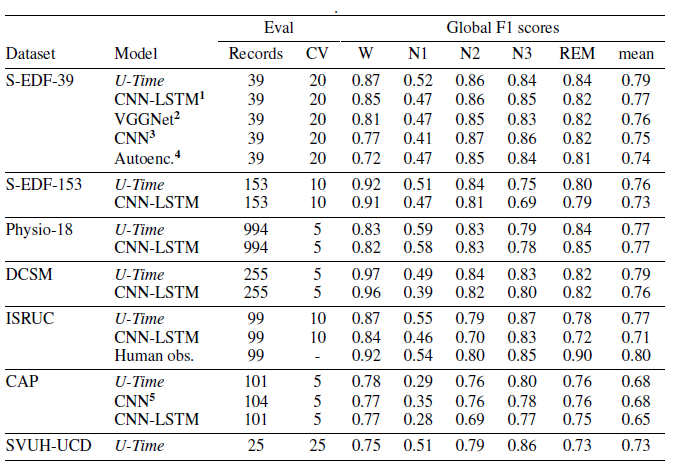
U-net was applied to 7 different PSG datasets with fixed architecture and hyperparameters, so there was no data-specific tuning. Furthermore, U-net only received one EEG channel as input.
The performance of U-net was compared to known models trained for use on a specific data set where available. As a baseline measure, the authors use an improved version of DeepSleepNet, which employs convolutional and recurrent layers and was designed to be applicable to different data sets. In the table summarising the results, this model is denoted by CNN-LSTM (LSTM stands for long short term memory, an example of a recurrent architecture).
Across all datasets, U-net had a high performance score similar to or higher than any known state of the art automated method specifically designed for that data set and the baseline.
Conclusion
Across all seven different PSG datasets, the same U-Time network architecture and hyperparameter settings were used. The benefits of this are such that one can avoid parameter overfitting this way, and robustness from U-Time's fully convolutional, feed-forward only architecture allows for it to be readily used by non-experts across health-related disciplines. The U-Time network architecture also has desirable properties such as computational efficiency, flexibility for the input window to be dynamically adjusted (i.e. an entire PSG record can be scored in a single pass), and high temporal resolution in the sleep stage output. While the authors chose to consider only a single EEG channel, it would be of interest to have U-Time receive multiple input channels for sleep staging, including EOG (eye movement) which often provides important information for distinguishing between wake and REM sleep stages. Overall, U-Time network architecture is a robust and efficient approach for time series segmentation that can be implemented with ease by health and computational researchers (Perslev et al., 2019).
References
Bai, S., Kolter, J.Z., & Koltun, V. (2018). An empirical evaluation of generic convolutional and recurrent networks for sequence modeling. CoRR, abs/1803.01271.
Chen, Q. & Wu, R. (2017). CNN is all you need. CoRR, abs/1712.09662.
Claude, R., Guilpin, C., & Limoge, A. (1998). Review of neural network applications in sleep research. Journal of Neuroscience Methods, 79, 187-193.
Faust, O., Hagiwara, Y., Hong, T.J., Lih, O.S., & Acharya, U.R. (2018). Deep learning for healthcare applications based on physiological signals: A review. Computer Methods and Programs in Biomedicine, 161, 1-13.
Perslev, M., Darkner, S., Jensen, M.H., Jennum, P.J., 7 Igel, C. (2019). U-Time: A Fully Convolutional Network for Time Series Segmentation Applied to Sleep Staging. Department of Computer Science, University of Copenhagen.
Vaswani, A. Shazeer, N., Parmar, N., Uszkoreit, J., Jones, L., Gomez, A.N., Kaiser, L., and Polosukhin, I. (2017). Attention is all you need. CoRR, abs/1706.03762.
Yu, F., & Koltun, V. (2015). Multi-scale context aggregation by dilated convolutions. CoRR, abs/1511.07122.