Deep Learning for Cardiologist-level Myocardial Infarction Detection in Electrocardiograms
Presented by
Zihui (Betty) Qin, Wenqi (Maggie) Zhao, Muyuan Yang, Amartya (Marty) Mukherjee
Introduction
This paper presents an approach to detecting heart disease from ECG signals by fine-tuning the deep learning neural network, ConvNetQuake. For context, ConvNetQuake is a convolutional neural network that has been used in the past for Earthquake detection and location from a single waveform [4]. A deep learning approach was used due to the model's ability to be trained using multiple GPUs and terabyte-sized datasets. This, in turn, creates a model that is robust against noise. The purpose of this paper is to provide detailed analyses of the contributions of the ECG leads on identifying heart disease, to show the use of multiple channels in ConvNetQuake enhances prediction accuracy, and to show that feature engineering is not necessary for any of the training, validation, or testing processes. In this area, the combination of data fusion and machine learning techniques exhibits great promise to healthcare innovation, and the analyses in this paper help further this realization. The benefits of translating knowledge between deep learning and its real-world applications in health are also illustrated.
Previous Work and Motivation
The database used in previous works is the Physikalisch-Technische Bundesanstalt (PTB) database, which consists of ECG records. Previous papers used techniques, such as CNN, SVM, K-nearest neighbors, naïve Bayes classification, and ANN. From these instances, the paper observes several faults in the previous papers. The first being the issue that most papers use feature selection on the raw ECG data before training the model. Dabanloo and Attarodi [2] used various techniques such as ANN, K-nearest neighbors, and Naïve Bayes. However, they extracted two features, the T-wave integral and the total integral, to aid in localizing and detecting heart disease. Sharma and Sunkaria [3] used SVM and K-nearest neighbors as their classifier, but extracted various features using stationary wavelet transforms to decompose the ECG signal into sub-bands. The second issue is that papers that do not use feature selection would arbitrarily pick ECG leads for classification without rationale. For example, Liu et al. [1] used a deep CNN that uses 3 seconds of ECG signal from lead II at a time as input. The decision for using lead II compared to the other leads was not explained.
The issue with feature selection is that it can be time-consuming and impractical with large volumes of data. The second issue with the arbitrary selection of leads is that it does not offer insight into why the lead was chosen and the contributions of each lead in the identification of heart disease. Thus, this paper addresses these two issues through implementing a deep learning model that does not rely on feature selection of ECG data and to quantify the contributions of each ECG and Frank lead in identifying heart disease.
Model Architecture
The dataset, which was used to train, validate, and test the neural network models, consists of 549 ECG records taken from 290 unique patients. Each ECG record has a mean length of over 100 seconds.
This Deep Neural Network model was created by modifying the ConvNetQuake model by adding 1D batch normalization layers.
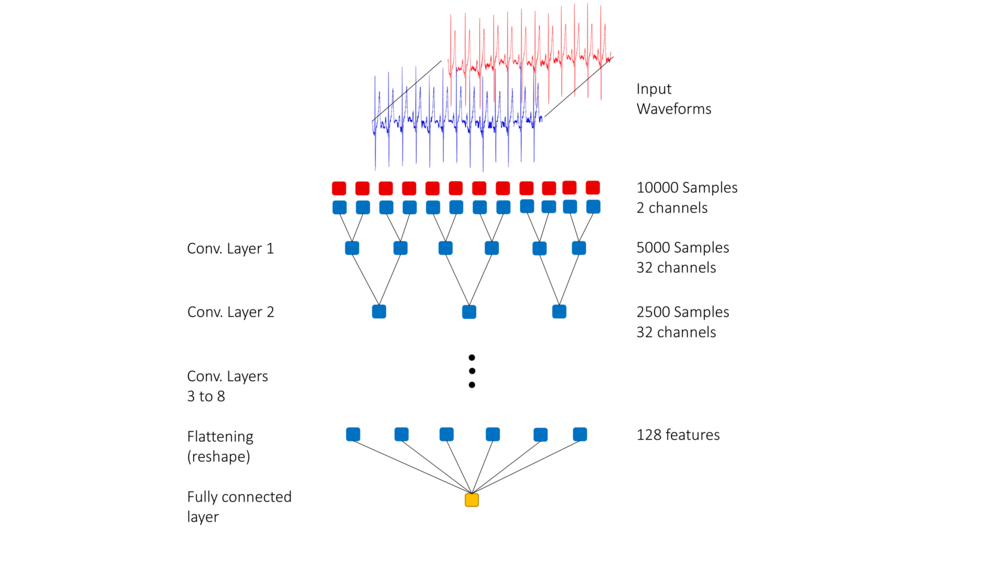
During the training stage, a 10-second long two-channel input was fed into the neural network. In order to ensure that the two channels were weighted equally, both channels were normalized. Besides, time invariance was incorporated by selecting the 10-second long segment randomly from the entire signal.
The input layer is a 10-second long ECG signal. There are 8 hidden layers in this model, each of which consists of a 1D convolution layer with the ReLu activation function followed by a batch normalization layer. The output layer is a one-dimensional layer that uses the Sigmoid activation function.
This model is trained by using batches of size 10. The learning rate is 10^-4. The ADAM optimizer is used. In training the model, the dataset is split into a train set, validation set, and test set with ratios 80-10-10.
During the training process, the model was trained from scratch numerous times to avoid inserting unintended variation into the model by randomly initializing weights.
Result
The paper first uses quantification of accuracies for single channels with 20-fold cross-validation, resulting in the highest individual accuracies: v5, v6, vx, vz, and ii. The researcher further investigated the accuracies for pairs of the top 5 highest individual channels using 20-fold cross-validation. The arrived at the conclusion of highest pairs accuracies to fed into a neural network is lead v6 and lead vz. They then use 100-fold cross validation on v6 and vz pair of channels, then compare outliers based on top 20, top 50 and total 100 performing models, finding that standard deviation is non-trivial and there are few models performed very poorly.
Next, they discussed 2 factors affecting model performance evaluation: 1) Random train-val-test split might have effects on the performance of the model, but it can be improved by access with a larger data set and further discussion; and 2) random initialization of the weights of the neural network shows little effects on the performance of the model performance evaluation, because of showing high average results with a fixed train-val-test split.
Comparing with other models in the other 12 papers, the model in this article has the highest accuracy, specificity, and precision. With concerns of patients' records affecting the training accuracy, they used 290 fold patient-wise split, resulting in the same highest accuracy of the pair v6 and vz same as record-wise split. Even though the patient-wise split might result in lower accuracy evaluation, however, it still maintains a high average of 97.83%.
Conclusion & Discussion
The paper introduced a new architecture for heart condition classification based on raw ECG signals using multiple leads. It outperformed the state-of-art model by a large margin of 1 percent. This study finds that out of the 15 ECG channels(12 conventional ECG leads and 3 Frank Leads), channel v6, vz, and ii contain the most meaningful information for detecting myocardial infraction. Also, recent advances in machine learning can be leveraged to produce a model capable of classifying myocardial infraction with a cardiologist-level success rate. To further improve the performance of the models, access to a larger labeled data set is needed. The PTB database is small. It is difficult to test the true robustness of the model with a relatively small test set. If a larger data set can be found to help correctly identify other heart conditions beyond myocardial infraction, the research group plans to share the deep learning models and develop an open-source, computationally efficient app that can be readily used by cardiologists.
A detailed analysis of the relative importance of each of the standard 15 ECG channels indicates that deep learning can identify myocardial infraction by processing only ten seconds of raw ECG data from the v6, vz and ii leads and reaches a cardiologist-level success rate. Deep learning algorithms may be readily used as commodity software. The neural network model that was originally designed to identify earthquakes may be re-designed and tuned to identify myocardial infraction. Feature engineering of ECG data is not required to identify myocardial infraction in the PTB database. This model only required ten seconds of raw ECG data to identify this heart condition with cardiologist-level performance. Access to a larger database should be provided to deep learning researchers so they can work on detecting different types of heart conditions. Deep learning researchers and the cardiology community can work together to develop deep learning algorithms that provide trustworthy, real-time information regarding heart conditions with minimal computational resources.
Fourier Transform(such as FFT) can be helpful when dealing with ECG signals. It transforms signals from time domain to frequency domain, which means some hidden features in frequency may be discovered.
Critiques
- The lack of large, labelled data sets is often a common problem in most applied deep learning studies. Since the PTB database is as small as you describe it to be, the robustness of the model which may be hard to gauge. There are very likely various other physical factors that may play a role in the study which the deep neural network may not be able to adjust for as well, since health data can be somewhat subjective at times and/or may be somewhat inaccurate, especially if machines are used to measurement. This might mean error was propagated forward in the study.
- Additionally, there is a risk of confirmation bias, which may occur when a model is self-training, especially given the fact that the training set is small.
- I feel that the results of deep learning models in medical settings where the consequences of misclassification can be severe should be evaluated by assigning weights to classification. In case if the misclassification can lead to severe consequences, then the network should be trained in such a way that it errs towards safety. For example, in case if heart disease, the consequences will be very high if the system says that there is no heart disease when in fact there is. So, the evaluation metric must be selected carefully.
- This is a useful and meaningful application topic in machine learning. Using Deep Learning to detect heart disease can be very helpful if it is difficult to detect disease by looking at ECG by humans eys. This model also useful for doing statistics, such as calculating the percentage of people get heart disease. But I think the doctor should not 100% trust the result from the model, it is almost impossible to get 100% accuracy from a model. So, I think double-checking by human eyes is necessary if the result is weird. What is more, I think it will be interesting to discuss more applications in mediccal by using this method, such as detecting the Brainwave diagram to predict a person's mood and to diagnose mental diseases.
- Compared to the dataset for other topics such as object recognition, the PTB database is pretty small with only 549 ECG records. And these are highly unbiased(Table 1) with 4 records for myocarditis and 148 for myocardial infarction. Medical datasets can only be labeled by specialists. This is why these datasets are related small. It would be great if there will be a larger, more comprehensive dataset.
References
[1] Na Liu et al. "A Simple and Effective Method for Detecting Myocardial Infarction Based on Deep Convolutional Neural Network". In: Journal of Medical Imaging and Health Informatics (Sept. 2018). doi: 10.1166/jmihi.2018.2463.
[2] Naser Safdarian, N.J. Dabanloo, and Gholamreza Attarodi. "A New Pattern Recognition Method for Detection and Localization of Myocardial Infarction Using T-Wave Integral and Total Integral as Extracted Features from One Cycle of ECG Signal". In: J. Biomedical Science and Engineering (Aug. 2014). doi: http://dx.doi.org/10.4236/jbise.2014.710081.
[3] L.D. Sharma and R.K. Sunkaria. "Inferior myocardial infarction detection using stationary wavelet transform and machine learning approach." In: Signal, Image and Video Processing (July 2017). doi: https://doi.org/10.1007/s11760-017-1146-z.
[4] Perol Thibaut, Gharbi Michaël, and Denolle Marin. "Convolutional neural network for earthquake detection and location". In: Science Advances (Feb. 2018). doi: 10.1126/sciadv.1700578