SPINS Tutorial: Difference between revisions
(Complete the SPINS tutorial page) |
(Add description of parameters) |
||
| Line 69: | Line 69: | ||
restart_from_dump = true | restart_from_dump = true | ||
== | == Change == | ||
If after analyzing the simulation you realize that you wanted to change a parameter (it wasn't quite right), then you only need to change the parameter in the configuration file (as opposed to older editions of SPINS where you'd need to recompile the entire code). | If after analyzing the simulation you realize that you wanted to change a parameter (it wasn't quite right), then you only need to change the parameter in the configuration file (as opposed to older editions of SPINS where you'd need to recompile the entire code). | ||
Here is a description of the problem parameters: | |||
--delta_rho arg density difference between upper and lower layers (as a percentage of rho_0) | |||
--pyc_loc arg Pycnocline location | |||
--h_halfwidth arg Pycnocline half-width | |||
--delta_x arg Horizontal transition half-width | |||
--H2 arg Height of mixed region (mode-2) - sets mode-2 wave amplitude | |||
--H1 arg Heigth of mixed region (mode-1) - sets mode-1 wave amplitude | |||
--L1 arg Length of region 1 (mode-2) | |||
--L2 arg Length of region 2 (mode-1) | |||
--dye1_loc arg Location of dye 1 | |||
--dye2_loc arg Location of dye 2 | |||
--dye1_width arg Width of dye 1 | |||
--dye2_width arg Width of dye 2 | |||
--dye_halfwidth arg Sharpness of the dye transition | |||
--enable_tracer Enable evolution of a second tracer | |||
If you run the simulation in the same directory, remember to clear all simulation files (output and diagnostic files). | If you run the simulation in the same directory, remember to clear all simulation files (output and diagnostic files). | ||
Revision as of 20:01, 5 March 2019
This is a tutorial for getting up to speed with SPINS using the case mode1_mode2. You must first have followed the instructions to install SPINS and compile the case file.
Run
Transfer the executable (mode1_mode2.x) and the configuration file (spins.conf) into a clean directory.
Run simulation by executing mpirun -np 4 ./mode1_mode2.x on the command line, or submit the job to the queue on a Compute Canada system with a submit script (see submitting jobs to Graham).
This will create two types of files
- Output files
- These are of the form
rho.3where the name (rho) is the variable and the extension is the output number
- - For example rho.3 is the density field at output number 3. Zero-indexing is used, so this is the 4th density file
- Since these can be memory expensive, they are created relatively infrequently
- These are of the form
- Diagnostic files
- These will generally be text files and contain information about the simulation
| Diagnostic Files | |||
|---|---|---|---|
| File | Optional | Matlab plotter | Description |
| diagnostics.txt | No | plot_diagnos | High temporal resolution information about the simulation (occurs at every time step). Examples include: KE, max velocity, energy budget terms, simulation timings. |
| stresses.txt (or stresses_top.txt and/or stresses_bottom.txt) | Yes | plot_stress | Surface forces (yes, it's a misnomer and needs to be fixed) at the top/bottom surfaces. Only useful if no slip boundary condition is used. Multiple integrals along the surfaces are completed. |
| plot_times.txt | No | plot_diagnos | Explicit statement of the time associated with each output. Also gives the time to write output files. |
Analyze
Matlab and Python are the primary languages used for analyzing SPINS simulations. These are great for computation of quantities specific to your study. The Python (SPINSpy) and Matlab (SPINSmatlab) packages contain much of the important tools for each language. Three dimensional visualization should use Paraview or VisIt (see Visualization).
Here, we will use Matlab to make a simple plot of the density field. We assume that the SPINSmatlab package is on your path (as is the case for boogaloo).
gdpar = spins_gridparams(); split_gdpar- This reads in the grid and parameters and places them in a structure (
gdpar).split_gdparseparates the two sub-structures into the structuresparandgd, which are the parameters and grid, respectively. parcontains information fromspins.confin addition to further simulation parametersgdcontains the grid. x,y,z grids can be extracted withgd.x,gd.y,orgd.z- The default setting is that the grid is unmapped and vector grids are produced, to get the full grid use
gdpar = spins_gridparams('FastFull');orgdpar = spins_gridparams('Full');
- This reads in the grid and parameters and places them in a structure (
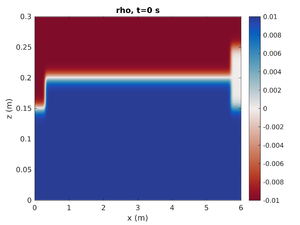
spins_plot2d('rho',0);- First argument is the field to plot
- Second argument is the output number. The simulation time can also be specified in this argument as a string.
spins_plot2d('rho','2');will plot the simulation at the closest output to t=2s. Many optional arguments exist (typehelp spins_plot2dfor more options).
To read an individual file:
u = spins_reader('u',10)
Restart
Should you need to restart the simulation (due to expenditure of allocated time, require more time, a node failure, or otherwise) simply change the restart flags in the configuration file.
The simplest change is to set
restart = true restart_time = 5.5 restart_sequence = 11
where the simulation will restart at output 11 which corresponds to t=5.5s. These numbers can be found as the last row in plot_time.txt.
SPINS also has an automatic safety dump if the allocated time is close to expiring (See automatically specify the runtime for more info). In this case, if the safety write was done successfully then you only need to set
restart_from_dump = true
Change
If after analyzing the simulation you realize that you wanted to change a parameter (it wasn't quite right), then you only need to change the parameter in the configuration file (as opposed to older editions of SPINS where you'd need to recompile the entire code).
Here is a description of the problem parameters:
--delta_rho arg density difference between upper and lower layers (as a percentage of rho_0) --pyc_loc arg Pycnocline location --h_halfwidth arg Pycnocline half-width --delta_x arg Horizontal transition half-width --H2 arg Height of mixed region (mode-2) - sets mode-2 wave amplitude --H1 arg Heigth of mixed region (mode-1) - sets mode-1 wave amplitude --L1 arg Length of region 1 (mode-2) --L2 arg Length of region 2 (mode-1) --dye1_loc arg Location of dye 1 --dye2_loc arg Location of dye 2 --dye1_width arg Width of dye 1 --dye2_width arg Width of dye 2 --dye_halfwidth arg Sharpness of the dye transition --enable_tracer Enable evolution of a second tracer
If you run the simulation in the same directory, remember to clear all simulation files (output and diagnostic files).
Only Mode-1 ISW
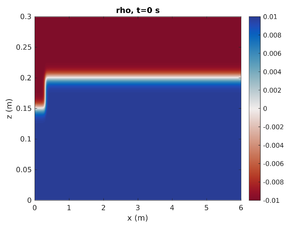
To change the simulation to only have the left mode-1 ISW change spins.conf to have:
H2 = 0.0
This will set the mode-2 amplitude to zero.
Only Mode-2 ISW
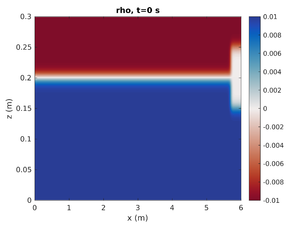
To change the simulation to only have the right mode-2 ISW change spins.conf to have:
H1 = 0.0
This will set the mode-1 amplitude to zero.
Gravity Current
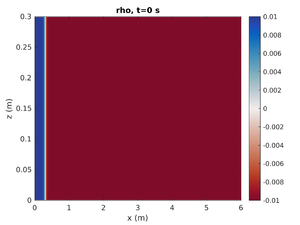
To change the simulation to have a rightward moving gravity current change spins.conf to have:
pyc_loc = -0.2 H1 = -1.0 H2 = 0.0
This will move the pycnocline below the bottom boundary, set the mode-1 amplitude to result in a region of high density at on end of the tank, and give no mode-2 contribution.
Try it yourself. See what each parameter does when they're changed.